 |
 |
- Search
| Clin Exp Reprod Med > Volume 47(3); 2020 > Article |
|
Abstract
Objective
The aim of this study was to investigate microRNAs (miRNAs) related to follicle-stimulating hormone (FSH) responsiveness using miRNA microarrays and to identify their target genes to determine the molecular regulatory pathways involved in FSH signaling in KGN cells.
Methods
To change the cellular responsiveness to FSH, KGN cells were treated with FSH receptor (FSHR)-specific small interfering RNA (siRNA) followed by FSH. miRNA expression profiles were determined through miRNA microarray analysis. Potential target genes of selected miRNAs were predicted using bioinformatics tools, and their regulatory function was confirmed in KGN cells.
Results
We found that six miRNAs (miR-1261, miR-130a-3p, miR-329-3p, miR-185-5p, miR-144-5p and miR-4463) were differentially expressed after FSHR siRNA treatment in KGN cells. Through a bioinformatics analysis, we showed that these miRNAs were predicted to regulate a large number of genes, which we narrowed down to cytochrome P450 family 19 subfamily A member 1 (CYP19A1) and estrogen receptor alpha (ESR1) as the main targets for miR-4463. Functional analysis revealed that miR-4463 is a regulatory factor for aromatase expression and function in KGN cells.
Conclusion
In this study, we identified differentially expressed miRNAs related to FSH responsiveness. In particular, upregulation of miR-4463 expression by FSHR deficiency in human granulosa cells impaired 17β-estradiol synthesis by targeting CYP19A1 and ESR1. Therefore, our data might provide novel candidates for molecular biomarkers for use in research into poor responders.
Controlled ovarian hyperstimulation (COH) is one of the main stages of assisted reproductive technology (ART) treatment. Women who show suboptimal responsiveness to exogenous gonadotropins during COH are referred to as poor responders [1]. Poor responders typically have low in vitro fertilization (IVF) success rates or experience cycle termination because few oocytes are received. Accordingly, they often need prolonged ovarian stimulation to achieve an appropriate ovarian response and to obtain enough oocytes for IVF [2]. Poor outcomes during IVF cycles in poor responders can cause tremendous emotional stress and increase the financial burden for the patient, which can result in them abandoning IVF treatment. Thus, improved strategies for ovarian stimulation in poor responders are essential for maximizing the success rate of IVF.
Although ovarian reserve tests are performed to predict responses and outcomes prior to IVF, they have little clinical value, and gonadotropin dosing is often decided based on empirical evidence [3]. In addition, several gene polymorphisms in the follicle-stimulating hormone (FSH) receptor (FSHR), estrogen receptor (ER), and luteinizing hormone receptor also seem to be important factors in determining the ovarian response in COH [4]. However, to date, no parameter can accurately predict the ovarian response to COH. The molecular mechanisms responsible for poor response to COH remain largely unknown.
In this study, we focused on FSHR dysfunction as a cause of poor ovarian response (POR). In a previous study, female FSHR+/− mice showed reduced fertility with profound oocyte loss and low 17β-estradiol levels, which are similar features to those observed in poor responders [5]. Indeed, the protein level of FSHR was found to be lower in granulosa cells (GCs) of poor responders than in GCs of moderate and high responders and was positively correlated with the peak level of serum 17β-estradiol and the number of mature oocytes [6]. In addition, among the several identified polymorphisms of the FSHR gene, two (Ser680Asn and Thr307Ala) are the most commonly investigated [7]. These polymorphisms have been found to influence the ovarian response to FSH stimulation and are also associated with polycystic ovary syndrome (PCOS) [7,8].
Due to the size of microRNAs (miRNAs; 19-25 bp), they can be easily detected by polymerase chain reaction (PCR). Therefore, miRNA is an attractive target for use as a predictor. Mature miRNA binds to 3'-untranslated regions (UTRs) of target mRNA and causes the degradation of mRNA and translational repression, exhibiting diverse functions in the regulation of cell growth, differentiation, survival, and apoptosis [9,10]. Knockout of Dicer, which is the ribonuclease III that is involved in the maturation of miRNAs, in the mouse ovary causes dysfunctions in folliculogenesis, oocyte maturation, ovulation, and fertility [11], suggesting the significance of miRNA for ovarian function. Steroidogenesis-related miRNAs, such as miR-133b, miR-378 and miR-224, regulate 17β-estradiol synthesis and release [12-14]. miR-15a and miR-188 participate in controlling human ovarian cell proliferation and apoptosis [15]. Furthermore, miR-23a, which targets XIAP, is expressed in patients with premature ovarian failure and plays a role in GC apoptosis [16]. However, little is known about the regulatory roles of miRNAs in the ovarian response to gonadotropins.
It has been reported that GC dysfunction is associated with infertility. In women with PCOS, morphologically abnormal GCs that have significantly low levels of estrogen and progesterone receptors contribute to the arrest of follicle development and to an increase in reactive oxygen species generation that affects the IVF success rate [17,18]. Compared to GCs from moderate and high responders, GCs obtained from poor responders showed a significantly lower expression level of FSHR [6]. Thus, studies on the physiological function of GCs are required to understand the mechanisms that underlie the development of various compartments of growing ovarian follicles, thereby yielding insights into therapeutic strategies for PCOS and POR. Although human GCs are easily isolated, they are only obtainable in small numbers and cannot survive in culture medium for extended cell generations due to their luteinizing properties. To overcome these limitations, immortalized human granulosa-like cell lines have been established for many years [19]. Among them, we used KGN cells with FSH-sensitive aromatase activity to develop an in vitro model that mimicked the GC status of poor responders.
The aims of this study were to investigate differentially expressed miRNAs that depend on the FSH responsiveness of GCs, to identify new potential miRNAs as predictors of POR, and to elucidate the molecular mechanism of POR.
A KGN immortalized human GC line was obtained from Riken BioResource Center (Riken Cell Bank, Tsukuba, Japan) [20]. The cells were maintained in a 1:1 mixture of DMEM and Ham’s F-12 medium (Gibco, Grand Island, NY, USA) containing 10% fetal bovine serum (FBS). Human SKOV3 ovarian cancer cells were obtained from the Korean Cell Line Bank (KCLB, Seoul, Korea). The cells were maintained in RPMI-1640 medium (Gibco) containing 10% FBS, 25 mM HEPES (Gibco), 25 mM NaHCO3, and 1% penicillin/streptomycin (Gibco). Primary human ovarian surface epithelial (HOSE) cells were purchased from Innoprot (Derio, Spain) and were cultivated in ovarian surface epithelial cell medium (Innoprot), which is a complete medium designed for the optimal growth of normal human ovarian epithelial cells in vitro. For the adherence of HOSE cells, 1 mg/mL of poly-L-lysine stock solution was added to culture dishes at the concentration of 2 μg/cm2, and the culture dishes were left in a 37°C incubator overnight.
FSHR small interfering RNA (siRNA) and nontargeting siRNA were purchased from Dharmacon (Lafayette, CO, USA). One day before transfection, KGN cells were seeded in six-well plates and transfected on the next day at 70% confluence. The cells were then transfected with 50 nM FSHR siRNA or nontargeting siRNA for 48 hours using Lipofectamine (Invitrogen, Carlsbad, CA, USA) followed by FSH (1 IU/mL; LG Chem, Seoul, Korea) treatment for an additional 24-hour period. At 72 hours after the treatment with siRNA and FSH, the cells were harvested for further analysis.
miRNA expression was analyzed with a miRCURY LNA microRNA Array (7th gen-has, mmu & rno array; Exiqon, Vedbaek, Denmark) covering 690 well-characterized rat miRNAs among 3,100 capture probes for human, mouse, and rat miRNAs. In this procedure, 5’-phosphates from 250–1,000 ng of total RNA were removed by calf intestinal alkaline phosphatase treatment, followed by labeling with Cy3 green fluorescent dye. The labeled samples were subsequently hybridized by loading onto a microarray slide using a Hybridization Chamber Kit (part #G2534A; Agilent Technologies, Santa Clara, CA, USA) and a Hybridization Gasket Slide Kit (part #G2534-60003; Agilent Technologies). Hybridization was performed over 16 hours at 56°C, followed by washing the microarray slide as recommended by the manufacturer. The processed microarray slides were then scanned with an Agilent G2565CA Microarray Scanner System (Agilent Technologies). Scanned images were imported using Agilent Feature Extraction software version 10.7.3.1 (Agilent Technologies), and the fluorescence intensities of each image were quantified using a modified version of the Exiqon protocol and corresponding gene array list files.
The data were processed based on a quantile normalization method using GeneSpring GX 13.1 (Agilent Technologies). This normalization method aims to make the distribution of intensities for each array in a set of arrays the same. The normalized and log-transformed intensity values were then analyzed using GeneSpring GX 13.1 (Agilent Technologies).
The target genes of the miRNAs (miR-130a-3p, miR-185-5p, miR329-3p and miR-4463) were predicted using eight distinct in silico target prediction tools: miRWalk (http://mirwalk.umm.uni-heidelberg.de/), miRanda (http://www.microrna.org/microrna/home.do), miRDB (http://www.mirdb.org/), Pictar2 (https://pictar.mdc-berlin.de/), PITA (https://genie.weizmann.ac.il/pubs/mir07/mir07_prediction.html), RNA22 (https://cm.jefferson.edu/rna22/Interactive/), RNAhybrid (https://bibiserv.cebitec.uni-bielefeld.de/rnahybrid), and TargetScan (http://www.targetscan.org/vert_72/), with a p-value threshold of 0.05. Functional pathway enrichment analysis of the target genes was conducted using the Database for Annotation, Visualization and Integrated Discovery (DAVID) online database (https://david.ncifcrf.gov/).
Total RNA was extracted using an miRNeasy Micro Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. For gene expression analysis, RNA (2 μg) was reverse-transcribed into first-strand cDNA with oligo (dT) and Moloney murine leukemia virus reverse transcriptase (Promega, Madison, WI, USA). Quantitative realtime PCR (qRT-PCR) analysis was performed using an iCycler (Bio-Rad, Hercules, CA, USA), iQ SYBR Green Supermix PCR reagents (Bio-Rad) were used for monitoring the amplification, and the results were evaluated with the iCycler iQ real-time detection system software. The amplification mixture contained cDNA, 5 pmol of forward and reverse primers, and SYBR Green Supermix. Templates were amplified with 40 cycles of denaturation at 95°C for 40 seconds, annealing at 60°C for 40 seconds, and extension at 72°C for 40 seconds. Upon the completion of PCR, the fluorescence was monitored while slowly heating the samples from 55°C to 95°C at 0.5°C intervals. Human GAPDH was used as the endogenous reference for mRNA normalization.
For miRNA expression analysis, an HB miR multi assay kit system I (HeimBiotek, Seongnam, Korea) was used for the analysis of miRNA in a two-step process. First, RNA (500 ng) was reverse-transcribed using an HB_I RT Reaction Kit and its reagents at 37°C for 60 minutes, followed by incubation at 95°C for 5 minutes for one cycle. Second, an HB_I Real-Time PCR Master Mix Kit was used to amplify the synthesized cDNAs with an iCycler (Bio-Rad). qRT-PCR involved an initial activation for 15 minutes at 95°C, 45 thermal cycles at 95°C for 10 seconds and 60°C for 40 seconds and standard melting conditions of a ramp ranging from 55°C to 95°C with a 0.5°C increase for each step. The primers used for miRNA amplification were purchased from HeimBiotek, and RNU6B was used as the endogenous reference for miRNA normalization. After mRNA or miRNA normalization, fold changes were calculated using the 2–ΔΔt method. All experiments were conducted in triplicate and repeated at least three times.
A miR-4463 mimic and inhibitor were purchased from Bioneer (Daejeon, Korea). KGN cells were transfected with 10 nM miR-4463 mimic or 50 nM miR-4463 inhibitor for 48 hours using Lipofectamine 2000 (Invitrogen).
Cellular proteins were extracted from treated cells and homogenized in radioimmunoprecipitation assay lysis buffer with a 1% protease inhibitor cocktail (Thermo Scientific, Waltham, MA, USA). Protein extracts (30 μg) were separated using sodium dodecyl sulfate polyacrylamide gel electrophoresis and then transferred onto polyvinylidene difluoride membranes. The membranes were blocked for 1 hour in Tris-buffered saline/Tween (TBST; 0.2 M NaCl, 0.1% Tween-20, and 10 mM Tris [pH 7.4]) containing 5% nonfat dry milk. The membranes were incubated overnight at 4°C with diluted primary antibodies against FSHR (ENT1795; Elabscience, Wuhan, China), ERα (sc542; Santa Cruz Biotechnology, Santa Cruz, CA, USA), ERβ (sc-8974, Santa Cruz Biotechnology), CYP19A1 (3599-100; BioVision, Mountain View, CA, USA) and β-actin (PA1-183, Invitrogen). β-Actin was used as an internal control. The membranes were then washed several times with TBST and incubated with diluted secondary antibodies for 1 hour at room temperature. After washing, signals were developed using an enhanced chemiluminescence system (Amersham Biosciences, Piscataway, NJ, USA), and the relative expression of the protein bands was quantified using a chemiDoc XRS+ imaging system with Image Lab software.
Concentrations of 17β-estradiol secreted from KGN cells were determined by a competitive-based enzyme-linked immunosorbent assay (ELISA) using a human estrogen ELISA kit (MyBioSource, San Diego, CA, USA). KGN cells were seeded into six-well plates and transfected with siRNAs or the miRNA mimic. After incubation for 48–72 hours, the culture medium was collected to measure the 17β-estradiol levels secreted from the cultured KGN cells according to the manufacturer’s instructions. The absorbance was determined at a wavelength of 450 nm using an automated microplate reader (E-Max, Molecular Devices, Sunnyvale, CA, USA).
The data are presented as the mean ± standard error of the mean from at least three separate and independent experiments. Differences among multiple groups (mock, miR-4463 mimic, miR-4463 inhibitor, and miR-4463 mimic+FSH) were analyzed using one-way analysis of variance, followed by the Student-Newman-Keuls test. Differences between negative control and FSHR siRNA-treated groups were analyzed using the Student t-test. The statistical analysis was performed using MedCalc ver. 19.1.7 (MedCalc Software, Mariekerke, Belgium). The significance of differences (p<0.05) between the mean values of triplicate samples were determined for all experiments.
To establish an in vitro study using human GCs, we used KGN cells, which have similar physiological features to those of normal GCs, including functional FSHR expression and responsiveness to FSH [20]. Compared to the other widely used ovarian cell lines (HOSE and SKOV3, which are human ovarian carcinoma cells), KGN cells highly express FSHR both at the mRNA and protein levels, suggesting that KGN cells are useful tools for the in vitro study of human GCs (Figure 1). In addition, we examined the expression of two ERs, ERα and ERβ (encoded by ESR1 and ESR2, respectively), to verify the responsiveness to estrogens that GCs acquire for their rapid proliferation. Our findings showed that KGN cells expressed ESR1 and ESR2 mRNA at low levels, but the protein expression level of ERβ was greater than that of ERα (Figure 1). This finding is consistent with previous studies reporting that ERβ was more detectable than ERα in GCs, whereas ERα was more abundant in theca cells than in GCs in the ovary [21,22].
To mimic the GC status of poor responders with low expression levels of FSHR [6], KGN cells were transfected with FSHR siRNA for 72 hours to decrease the FSHR expression level and were treated with 1 U/mL FSH for 24 hours prior to the endpoint of siRNA transfection. Compared to negative control groups, the knockdown of FSHR by siRNA led to an approximately 30% reduction in mRNA and protein expression, and additional FSH treatment significantly increased the expression levels of FSHR at the mRNA and protein levels (Figure 2A and B). To evaluate 17β-estradiol levels after FSHR siRNA transfection and FSH treatment, the culture medium of transfected KGN cells was collected, and the 17β-estradiol levels were analyzed using ELISA. 17β-Estradiol levels were decreased by the knockdown of FSHR, and the subsequent FSH treatment after knockdown failed to restore the 17β-estradiol levels (Figure 2C). Overall, our observations demonstrated that additional FSH treatment after FSHR knockdown restored functional FSHR expression but failed to rescue the 17β-estradiol levels in KGN cells, suggesting that these cells are useful tools for the in vitro study of the physiological features of GCs in poor responders. These results suggest that it is difficult to recover the steroidogenic activity of human GCs in poor responders through additional FSH treatment following FSHR impairment.
To investigate differentially expressed miRNAs that depend on the status of FSHR expression, we profiled miRNA expression in the siFSHR+FSH (FSH treatment after FSHR siRNA transfection) group compared to the siControl+FSH (FSH treatment after control siRNA transfection) group. Fold changes ≥ 2 or ≤ 0.5 were considered to indicate differential expression. Among the six miRNAs, miR-185-5p showed increased expression, and miR-130a-3p, miR-1261, miR-144-5p, miR-329-3p, and miR-4463 showed decreased expression in the siFSHR+FSH group compared with that in the siControl+FSH group (Table 1). These six miRNAs were chosen to validate the microarray data using qRT-PCR analysis, which indicated that only miR-130a-3p showed the same trend as the data obtained from the microarray, whereas the remaining five miRNAs did not show consistent trends. In other words, miR-185-5p expression decreased and miR-329-3p and miR-4463 expression increased in the siFSHR+FSH group compared to the expression levels in the siControl+FSH group (Figure 3). miR-1261 and miR-144-5p were not detected in KGN cells by qRT-PCR analysis. Although the presence of false-positive results requires further discussion, it still worthwhile to note the identification of four differentially expressed miRNAs (miR-130a-3p, miR-185-5p, miR-329-3p and miR-4463) that are closely related to FSH responsiveness in human GCs. Further studies are required to understand their biological functions by predicting miRNA targets.
To determine the biological functions of the four differentially expressed miRNAs regulated by FSH signaling in GCs, the putative target genes of these miRNAs were identified using the following eight distinct in silico target prediction tools: miRWalk, miRanda, miRDB, Pictar2, PITA, RNA22, RNAhybrid, and TargetScan. These in silico approaches identified approximately 1,000–7,000 genes as targets for each miRNA. To narrow down the list of target genes, we classified these genes using two independent criteria. First, genes that were found in at least four of the eight bioinformatics sources were collected, according to which 1,658, 3,336, 1,707, and 4,256 genes were narrowed down as targets for miR-130a-3p, miR-185-5p, miR-329-3p, and miR-4463, respectively. Second, we sorted for genes that were regulated by at least two out of the four miRNAs selected from the microarray data. This workflow is shown in Figure 4. Finally, a total of 725 genes were selected for further analysis.
The DAVID online database was used to perform KEGG pathway enrichment analysis for the 725 selected target genes of the identified differentially expressed miRNAs. The top 10 significant pathways are listed and summarized in Table 2. Notably, these top 10 pathways include PI3K-AKT, RAS, and PKA/cAMP signaling, which are known to be significantly related to FSH-related signaling in GC differentiation and follicle growth [23-25]. These data strongly suggest that miR130a-3p, miR-185-5p, miR-329-3p, and miR-4463 are involved in FSH signaling in human GCs.
Based on the pathway enrichment analysis, we further analyzed the selected target genes of the four differentially expressed miRNAs. In the entire target gene list, FSH-related [24] and COH-related [26] genes were sorted, as shown in Table 3. Among them, we particularly focused on CYP19A1 and ESR1, which were found to be target genes of miR-4463, due to their correlation with steroidogenic activity in GCs. CYP19A1 and ESR1 were included among the target genes as FSH targets (CCND2, GSK3B, and CYP19A1) or COH-related genes (MTHFR, PGR, and ESR1). CYP19A1 encodes aromatase, which is the final enzyme for ovarian steroidogenesis, and ERα (encoded by ESR1) mediates the proliferative actions of estrogens [27,28]. This led us to further examine the regulation of miR-4463 in the steroidogenic activity of human GCs by targeting CYP19A1 and ESR1.
To investigate the association between FSHR and miR-4463 expression, we first observed changes in miR-4463 expression after FSHR suppression. It was found that miR-4463 expression time-dependently increased after transfection of FSHR siRNA (Figure 5A). We hypothesized that miR-4463 suppresses translation of CYP19A1 and ESR1 in KGN cells. A web tool (TargetScan) revealed that CYP19A1 and ESR1 are putative target genes of miR-4463, and putative miR-4463 binding sites in the 3'-UTRs of CYP19A1 and ESR1 were also identified (Figure 5B). To determine whether miR-4463 regulates the expression of CYP19A1 and ESR1, KGN cells were transfected with a miR4463 mimic or inhibitor to induce its functional overexpression or suppression, respectively. Immunoblotting analyses demonstrated that CYP19A1 protein expression levels decreased by approximately 40% after transfection with the miR-4463 mimic, whereas the miR4463 inhibitor treatment did not change the expression of CYP19A1. Although the ERα protein expression level was not affected by the miR-4463 mimic, it significantly increased by 22% with the inhibition of endogenous miR-4463 by the miR-4463 inhibitor treatment, indicating that miR-4463 also targets ESR1 (Figure 5C and D). Furthermore, ELISA analysis revealed that 17β-estradiol levels were significantly reduced by 30% in the miR-4463 mimic-transfected group, whereas treatment with the miR-4463 inhibitor tended to increase the 17β-estradiol levels compared with those in the mock control group (Figure 5E). Therefore, these data strongly suggest that miR4463 plays a crucial role in regulating 17β-estradiol synthesis by targeting CYP19A1 and ESR1 in KGN cells.
To further confirm that the inhibition of 17β-estradiol synthesis by miR-4463 is restored by FSH treatment, 1 U/mL FSH was administered for 24 hours after KGN cells were transfected with the miR-4483 mimic for 48 hours. Interestingly, the miR-4463-induced reductions in levels of CYP19A1 protein expression and 17β-estradiol were not restored by FSH treatment (Figure 5C-E). These data indicate that overexpression of miR-4463 precludes a role for FSH in 17β-estradiol synthesis in GCs with reduced expression of FSHR.
In the present study, we identified differentially expressed miRNAs in KGN cells mimicking the GCs of poor responders with low FSH responsiveness. These altered miRNAs are associated with FSH-related signaling pathways and are predicted to target several genes that function in the ovarian response. In particular, we found that miR4463 showed significantly increased expression in a state of low ovarian response and regulated 17β-estradiol synthesis by targeting CYP19A1 and ESR1. Our results showed that upregulation of miR4463 expression in poor responders might be one of the causes of POR with reduced 17β-estradiol production, suggesting that miR4463 is a potential predictor of POR.
Although the manifestations of POR are quite diverse, patients with POR are typically classified as having low ovarian reserve or normal ovarian reserve but an inherently low gonadotropin response [29,30]. Ovaries that respond poorly to hyperstimulation during IVF cycles induce the presence of a reduced number of FSH-sensitive follicles, which are most frequently associated with the condition of poor ovarian reserve, eventually leading to low success rates in pregnancy [31,32]. Because of individual variations between patients in the ovarian response, predicting the ovarian response to exogenous gonadotropin is the most important step in ovulation induction. Many parameters are used predict the ovarian response and to determine the starting dose of exogenous gonadotropin. For example, high levels of serum FSH [33], elevated levels of serum estradiol [34], decreased levels of serum inhibin B [33], and decreased levels of serum anti-Müllerian hormone [35] are measured in the early follicular phase. Sonographic findings, including decreased antral follicle count and decreased ovarian stromal blood flow, have also been suggested as predictors of ovarian response [36]. Despite the wide use of these predictors, a test that more easily and accurately predicts POR in patients undergoing IVF is required.
Numerous roles of differentially expressed miRNAs in various tissues of poor responders have been demonstrated in an increasing number of studies that have mainly focused on miRNAs as novel biomarkers of these ovarian conditions. Emerging evidence has demonstrated that miRNAs are involved in the modulation of gene expression and in a broad range of pathological conditions, including poor ovarian reserve [37]. Certain differentially expressed miRNAs in GCs of poor responders have been identified. These miRNAs target genes involved in signaling pathways associated with the ovarian response, such as the gonadotropin-releasing hormone signaling pathway, oocyte meiosis, the cell cycle, the insulin signaling pathway, and some cancer pathways [38]. miR-106a, which regulates cell viability and apoptosis of GCs via enhanced ASK1 signaling, has been reported to be downregulated in the serum and GCs of females with diminished ovarian reserve [39]. miR-21-5p, which is likely to be independent of serum estradiol levels, has been shown to be elevated in the cumulus cells of poor responders, suggesting that it may play an important role as a potential biomarker for oocyte or follicular viability [40]. The expression of miR-423-5p was significantly lower in hyper-responders than in normal responders. The predicted target genes of miR423-5p include CYP19A1, MTHFR, PGR, and FSHB, which are associated with the response to ovarian stimulation [38].
It has been reported that the functional expression of FSHR is an important indicator of POR, as aberrant FSHR activity has been found to lead to poor proliferation and differentiation of GCs and reduced production of 17β-estradiol [7]. FSHR polymorphisms have been reported to significantly affect the ovarian response to FSH stimulation [31,41]. This might imply that the FSH-FSHR axis plays a pivotal role in determining the status of GC growth and 17β-estradiol production at the center of physiological ovarian function.
GCs are indispensable for folliculogenesis and oocyte maturation in response to gonadotropin signaling. However, in vitro studies with GCs obtained from human ovaries using in vitro culture systems are impacted by the morphological changes of GCs into luteal cells, which lose proliferative activity and alter the gene profile [42]. To mimic the pathophysiological features of the GCs of poor responders in this study, we intentionally reduced FSHR gene expression by siRNA transfection to mimic the FORKO haploid animal model that has shown very similar physiological changes, such as low FSHR expression and low 17β-estradiol production in poor responders undergoing IVF [5]. Furthermore, the additional treatment with FSH after the knockdown of FSHR mimicked the administration of gonadotropin to poor responders in IVF cycles. Through these strategies, we identified differentially expressed miRNAs in poor responders under in vitro conditions.
In the present study, we identified four miRNAs (miR-130a-3p, miR-185-5p, miR-329-3p, and miR-4463) that were differentially expressed depending on FSH responsiveness in KGN cells. Most studies on miR-130a-3p, miR-185-5p, and miR-329-3p have been carried out in the field of cancer research. It has been reported that miR-130a-3p expression is downregulated in various cancers, including breast cancer, nasopharyngeal carcinoma, and esophageal squamous cell carcinoma, and that miR-130a-3p regulates cell proliferation, invasion, migration, the cell cycle, and the endothelial-mesenchymal transition [43-45]. Overexpression of miR-185-5p expression was found to promote apoptosis by reducing antiapoptotic BCL2L1 levels in prostate cancer cells [46]. In glioma, overexpression of the long non-coding RNA FOXD2–AS1 downregulated miR-185-5p expression following the upregulation of CCND2 and HMGA2 expression [47,48]. miR-329-3p was found to function as a tumor suppressor by directly targeting MAPK1 in cervical cancer [49]. Furthermore, a few reports have investigated the expression and functions of miR-4463 in vascular disorders, such as arteriosclerosis and carotid artery stenosis [50,51]. The upregulation of miR-4463 expression enhanced H2O2-induced oxidative stress and promotes apoptosis in endothelial cells [52]. The inhibition of miR-4463 promoted human umbilical vein endothelial cell survival and reduced apoptosis under hypoxic conditions by targeting the protein phosphatase-1 nuclear targeting subunit [53]. In addition, miR-4463 promoted vascular smooth muscle cell migration, which is implicated in vascular diseases, by targeting angiomotin [54]. Interestingly, a recent report has indicated that miR4463 expression was significantly upregulated in the serum of patients with PCOS, although its biological functions related to PCOS remain unknown [55].
The canonical pathway analysis in the present study using eight different bioinformatics tools identified 725 different target genes that were commonly regulated by miR-130a-3p, miR-185-5p, miR329-3p, and miR-4463. Among them, both CYP19A1 and ESR1 were predicted to be target genes of miR-4463, and we validated our prediction experimentally (Figure 5C and E). This result is notable because CYP19A1 and ESR1 are essential genes for 17β-estradiol production in ovarian GCs. In particular, CYP19A1 encodes an aromatase enzyme that plays a decisive role in the synthesis of 17β-estradiol in GCs from testosterone produced in theca cells [56]. miR-4463 mimics significantly reduced CYP19A1 protein levels and 17β-estradiol production in KGN cells, indicating that miR-4463 regulates 17β-estradiol synthesis via targeting CYP19A1. Based on a report that aromatase activity was reduced in PCOS follicles resulting from a genetic polymorphism of CYP19A1 [57], it is possible that the upregulation of miR-4463 expression in the serum of patients with PCOS [55] is associated with a poor response and low estrogen production. In order to clarify the existence of the FSHR/miR-4463/CYP19A1 axis in GCs, we additionally confirmed that suppression of FSHR decreased CYP19A1 levels in KGN cells (Figure 6). Furthermore, the failure to restore CYP19A1 expression and 17β-estradiol levels by FSH treatment after miR-4463 mimic transfection explains why subsequent FSH treatment after FSHR knockdown failed to restore 17β-estradiol levels despite the rescue of FSHR expression (Figure 2C). In other words, suppression of FSHR expression promotes miR-4463 expression, followed by impeding the action of FSH in GCs. These results imply that inhibition of miR-4463 expression must precede the improvement of responsiveness of GCs to FSH.
In summary, FSHR downregulation in poor responders causes overexpression of miR-4463 in GCs, followed by inhibition of translation of CYP19A1, leading to a decrease in the synthesis of 17β-estradiol (Figure 7). In this study, we first demonstrated that the elevation of miR-4463 expression by the attenuation of FSHR is a key feature that contributes to the low ovarian response in GCs. A comparison of miR4463 expression levels in the serum of poor responders and normal responders undergoing IVF is required to validate our results in cell lines. Further studies identifying signaling pathways relevant to the roles of miR-4463 in regulating 17β-estradiol synthesis in GCs of women with POR will expand our knowledge of the regulation of the ovarian response. Based on our findings, we suggest that miR-4463 is a new candidate for indicating individual differences in the response to exogenous gonadotropin. Therefore, monitoring changes in miR-4463 expression could be used to improve the accuracy of diagnoses for personalized ART treatment and to develop new markers for predicting the ovarian response in IVF cycles.
Figure 1.
The expression of follicle-stimulating hormone receptor (FSHR) and estrogen receptors (ERs) in KGN cells. Characterization of KGN cells by real-time polymerase chain reaction (A) and Western blot (B) for FSHR, which is a marker of granulosa cells, as well as ERα and ERβ. HOSE, human ovarian surface epithelial.
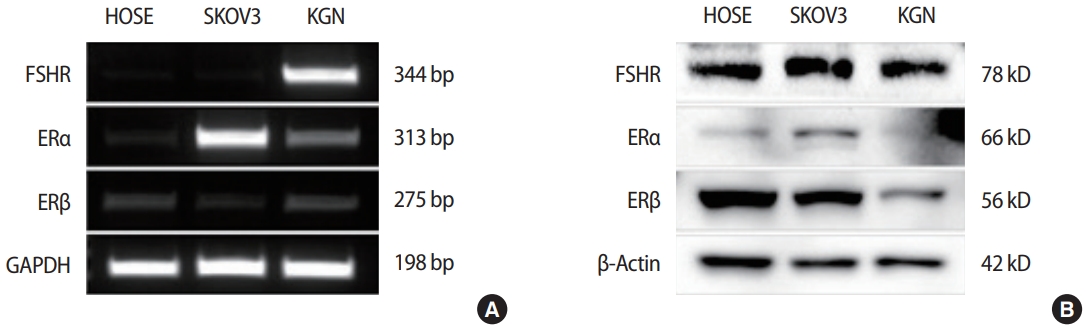
Figure 2.
Follicle-stimulating hormone (FSH) treatment restored the expression of FSH receptor (FSHR) but did not restore 17β-estradiol levels in FSHR small interfering RNA (siRNA)-transfected KGN cells. The levels of FSHR and 17β-estradiol were measured after KGN cells were cotreated with FSHR siRNA and FSH (1 U/mL). (A) The mRNA expression level of FSHR was calculated from CT values obtained by quantitative real-time polymerase chain reaction. The mRNA expression levels were normalized to the expression level of GAPDH, and relative expression was calculated. (B) The protein expression level of FSHR was measured by Western blot analysis. The relative protein expression levels were calculated by measuring the density and area of the bands. (C) 17β-Estradiol levels in the cell culture supernatant were measured by enzyme-linked immunosorbent assay. The dashed line on value 1 indicates the value of each negative control group. The experiments were repeated at least three times, and the data are presented as the mean ± standard error of the mean. siFSHR, FSHR siRNA; siFSHR+FSH, FSH treatment after FSHR siRNA transfection. a)p< 0.05, b)p< 0.001: indicate statistical significance.

Figure 3.
Verification of differential micro-RNA (miRNA) expression by quantitative real-time polymerase chain reaction (qRT-PCR). Fold differences determined by miRNA microarray and qRT-PCR analysis are shown on the Y-axis. The expression of each miRNA was normalized to that of RNU6B and was then evaluated using the 2–ΔΔt method, and the relative expression was calculated. Fold changes indicate the value of the siFSHR+FSH to siControl+FSH ratio. The dashed line on the value 1 indicates the value of each siControl+FSH group. qRTPCR analysis was biologically repeated three times. The error bars indicate the mean ± standard error of the mean. FSH, follicle-stimulating hormone; FSHR, FSH receptor; siFSHR, FSHR siRNA.
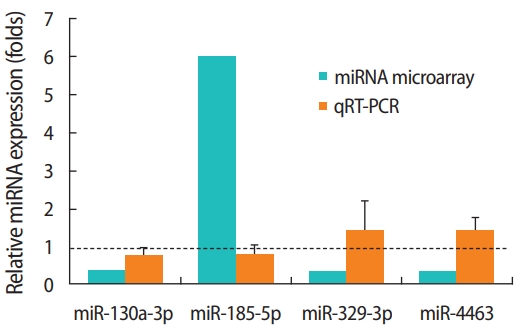
Figure 4.
The workflow for the selection of putative target genes of four differentially expressed miRNAs related to follicle-stimulating hormone signaling using eight bioinformatics tools. miRNA, micro-RNA.

Figure 5.
miR-4463 targets cytochrome P450 family 19 subfamily A member 1 (CYP19A1) and estrogen receptor alpha (ESR1) genes and inhibits steroidogenesis in KGN cells. After KGN cells were transfected with a miR-4463 mimic, miR-4463 inhibitor, and miR-4463 mimic with folliclestimulating hormone (FSH), the expression levels of CYP19A1, estrogen receptor α (ERα), and synthesis of 17β-estradiol were analyzed. (A) Expression of miR-4463 increased by knockdown of follicle-stimulating hormone receptor (FSHR) in KGN cells. (B) The putative sites in the 3’-untranslated region (UTR) of CYP19A1 and ESR1 mRNA contain the seed sequence for hsa-miR-4463 binding. The nucleotides in bold letters indicate the seed region of miR-4463. (C, D) Protein expression levels were measured by Western blot analysis, and β-actin was used as an internal control. The relative protein expression levels were calculated by measuring the density and area of the bands. (E) 17β-estradiol levels were measured by enzyme-linked immunosorbent assay using cell culture supernatant after transfection with miR-4463 mimic, miR-4463 inhibitor, and miR-4463 mimic with FSH. The experiments were repeated at least three times, and the data are presented as the mean ± standard error of the mean. siRNA, small interfering RNA. a)p< 0.05: indicate statistical significance. The p-values were obtained by one-way analysis of variance and the Student-Newman-Keuls test.
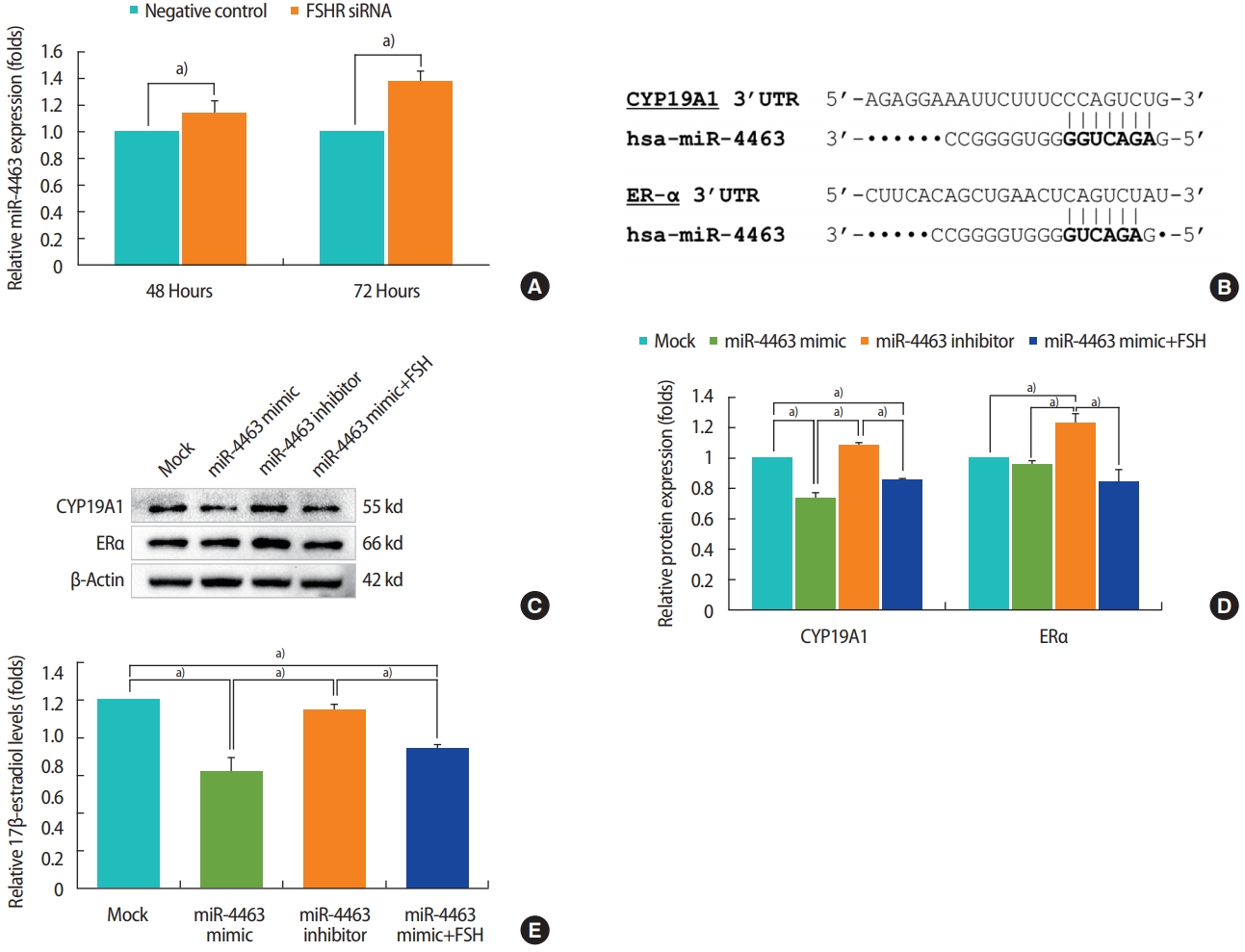
Figure 6.
Treatment of follicle-stimulating hormone receptor (FSHR) small interfering RNA (siRNA) decreased CYP19A1 expression. (A) The protein expression levels were measured by Western blot analysis, and β-actin was used as an internal control gene. (B) The relative protein expression levels were calculated by measuring the density and area of the bands. The experiments were repeated at least three times, and the data are presented as the mean ± standard error of the mean. NC, negative control group; siFSHR, FSHR siRNA-treated group. a)p< 0.001: indicates statistical significance.
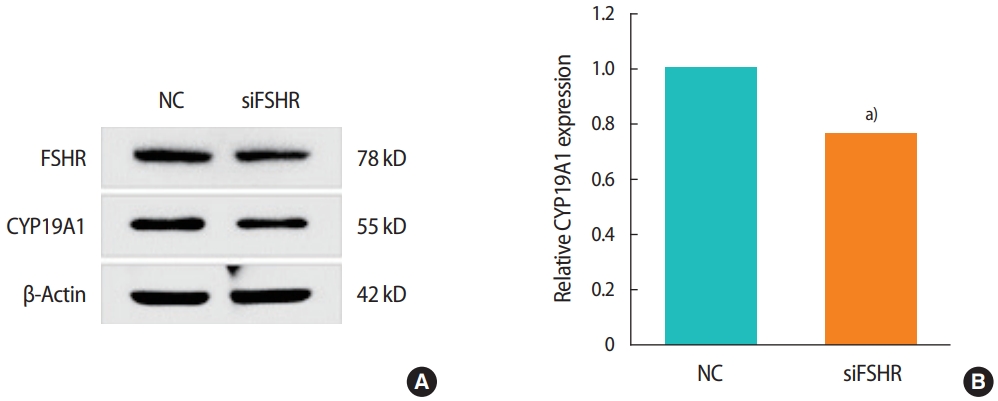
Figure 7.
The predicted mechanism of the regulatory role of miR-4463 on CYP19A1 in granulosa cells (GCs) for follicle-stimulating hormone-responsive 17β-estradiol synthesis. In this study, we suggested that follicle-stimulating hormone receptor (FSHR) downregulation in patients with poor responsiveness may increase miR-4463 expression levels in GCs, followed by the inhibition of CYP19A1 translation. Consequently, steroidogenesis, especially 17β-estradiol synthesis, is reduced.

Table 1.
Fold changes of differentially expressed miRNAs indicated by miRNA microarray and qRT-PCR validation results
| miRNA |
Fold change |
|
|---|---|---|
| miRNA microarray | qRT-PCR | |
| miR-130a-3p | 0.360 | 0.74 |
| miR-1261 | 0.329 | Not detected |
| miR-144-5p | 0.437 | Not detected |
| miR-185-5p | 5.938 | 0.77 |
| miR-329-3p | 0.312 | 1.38 |
| miR-4463 | 0.372 | 1.34 |
Table 2.
Top 10 KEGG pathways including the target genes of miR-130a-3p, miR-185-5p, miR-329-3p, and miR-4463
References
1. Tanbo T, Abyholm T, Bjoro T, Dale PO. Ovarian stimulation in previous failures from in-vitro fertilization: distinction of two groups of poor responders. Hum Reprod 1990;5:811-5.



2. Bar-Hava I, Yoeli R, Yulzari-Roll V, Ashkenazi J, Shalev J, Orvieto R. Controlled ovarian hyperstimulation: does prolonged stimulation justify cancellation of in vitro fertilization cycles? Gynecol Endocrinol 2005;21:232-4.


3. Fauser BC, Diedrich K, Devroey P, Evian Annual Reproduction Workshop Group 2007. Predictors of ovarian response: progress towards individualized treatment in ovulation induction and ovarian stimulation. Hum Reprod Update 2008;14:1-14.



4. Sheikhha MH, Eftekhar M, Kalantar SM. Investigating the association between polymorphism of follicle-stimulating hormone receptor gene and ovarian response in controlled ovarian hyperstimulation. J Hum Reprod Sci 2011;4:86-90.



5. Danilovich N, Maysinger D, Sairam MR. Perspectives on reproductive senescence and biological aging: studies in genetically altered follitropin receptor knockout [FORKO] mice. Exp Gerontol 2004;39:1669-78.


6. Cai J, Lou HY, Dong MY, Lu XE, Zhu YM, Gao HJ, et al. Poor ovarian response to gonadotropin stimulation is associated with low expression of follicle-stimulating hormone receptor in granulosa cells. Fertil Steril 2007;87:1350-6.


7. Simoni M, Gromoll J, Nieschlag E. The follicle-stimulating hormone receptor: biochemistry, molecular biology, physiology, and pathophysiology. Endocr Rev 1997;18:739-73.


8. Perez Mayorga M, Gromoll J, Behre HM, Gassner C, Nieschlag E, Simoni M. Ovarian response to follicle-stimulating hormone (FSH) stimulation depends on the FSH receptor genotype. J Clin Endocrinol Metab 2000;85:3365-9.



9. Stefani G, Slack F. MicroRNAs in search of a target. Cold Spring Harb Symp Quant Biol 2006;71:129-34.


11. Lei L, Jin S, Gonzalez G, Behringer RR, Woodruff TK. The regulatory role of Dicer in folliculogenesis in mice. Mol Cell Endocrinol 2010;315:63-73.


12. Yao G, Yin M, Lian J, Tian H, Liu L, Li X, et al. MicroRNA-224 is involved in transforming growth factor-beta-mediated mouse granulosa cell proliferation and granulosa cell function by targeting Smad4. Mol Endocrinol 2010;24:540-51.




13. Xu S, Linher-Melville K, Yang BB, Wu D, Li J. Micro-RNA378 (miR-378) regulates ovarian estradiol production by targeting aromatase. Endocrinology 2011;152:3941-51.




14. Dai A, Sun H, Fang T, Zhang Q, Wu S, Jiang Y, et al. MicroRNA-133b stimulates ovarian estradiol synthesis by targeting Foxl2. FEBS Lett 2013;587:2474-82.


15. Sirotkin AV, Laukova M, Ovcharenko D, Brenaut P, Mlyncek M. Identification of microRNAs controlling human ovarian cell proliferation and apoptosis. J Cell Physiol 2010;223:49-56.


16. Yang X, Zhou Y, Peng S, Wu L, Lin HY, Wang S, et al. Differentially expressed plasma microRNAs in premature ovarian failure patients and the potential regulatory function of mir-23a in granulosa cell apoptosis. Reproduction 2012;144:235-44.


17. Artimani T, Saidijam M, Aflatoonian R, Amiri I, Ashrafi M, Shabab N, et al. Estrogen and progesterone receptor subtype expression in granulosa cells from women with polycystic ovary syndrome. Gynecol Endocrinol 2015;31:379-83.


18. Karuputhula NB, Chattopadhyay R, Chakravarty B, Chaudhury K. Oxidative status in granulosa cells of infertile women undergoing IVF. Syst Biol Reprod Med 2013;59:91-8.


19. Havelock JC, Rainey WE, Carr BR. Ovarian granulosa cell lines. Mol Cell Endocrinol 2004;228:67-78.


20. Nishi Y, Yanase T, Mu Y, Oba K, Ichino I, Saito M, et al. Establishment and characterization of a steroidogenic human granulosalike tumor cell line, KGN, that expresses functional follicle-stimulating hormone receptor. Endocrinology 2001;142:437-45.



21. Nilsson S, Gustafsson JA. Biological role of estrogen and estrogen receptors. Crit Rev Biochem Mol Biol 2002;37:1-28.


22. Chu S, Nishi Y, Yanase T, Nawata H, Fuller PJ. Transrepression of estrogen receptor beta signaling by nuclear factor-kappab in ovarian granulosa cells. Mol Endocrinol 2004;18:1919-28.


23. Wayne CM, Fan HY, Cheng X, Richards JS. Follicle-stimulating hormone induces multiple signaling cascades: evidence that activation of Rous sarcoma oncogene, RAS, and the epidermal growth factor receptor are critical for granulosa cell differentiation. Mol Endocrinol 2007;21:1940-57.



24. Richards JS, Pangas SA. The ovary: basic biology and clinical implications. J Clin Invest 2010;120:963-72.



25. Gonzalez-Robayna IJ, Falender AE, Ochsner S, Firestone GL, Richards JS. Follicle-Stimulating hormone (FSH) stimulates phosphorylation and activation of protein kinase B (PKB/Akt) and serum and glucocorticoid-lnduced kinase (Sgk): evidence for A kinase-independent signaling by FSH in granulosa cells. Mol Endocrinol 2000;14:1283-300.


26. Altmae S, Hovatta O, Stavreus-Evers A, Salumets A. Genetic predictors of controlled ovarian hyperstimulation: where do we stand today? Hum Reprod Update 2011;17:813-28.



27. Ryan KJ. Biochemistry of aromatase: significance to female reproductive physiology. Cancer Res 1982;42:3342s-3344s.

29. Gerasimova T, Thanasoula MN, Zattas D, Seli E, Sakkas D, Lalioti MD. Identification and in vitro characterization of follicle stimulating hormone (FSH) receptor variants associated with abnormal ovarian response to FSH. J Clin Endocrinol Metab 2010;95:529-36.




30. Ferraretti AP, La Marca A, Fauser BC, Tarlatzis B, Nargund G, Gianaroli L, et al. ESHRE consensus on the definition of ‘poor response’ to ovarian stimulation for in vitro fertilization: the Bologna criteria. Hum Reprod 2011;26:1616-24.


31. Padhy N, Gupta S, Mahla A, Latha M, Varma T. Demographic characteristics and clinical profile of poor responders in IVF/ICSI: a comparative study. J Hum Reprod Sci 2010;3:91-4.



32. Hendriks DJ, te Velde ER, Looman CW, Bancsi LF, Broekmans FJ. Expected poor ovarian response in predicting cumulative pregnancy rates: a powerful tool. Reprod Biomed Online 2008;17:727-36.


33. Broekmans FJ, Kwee J, Hendriks DJ, Mol BW, Lambalk CB. A systematic review of tests predicting ovarian reserve and IVF outcome. Hum Reprod Update 2006;12:685-718.



34. Licciardi FL, Liu HC, Rosenwaks Z. Day 3 estradiol serum concentrations as prognosticators of ovarian stimulation response and pregnancy outcome in patients undergoing in vitro fertilization. Fertil Steril 1995;64:991-4.


35. Seifer DB, Maclaughlin DT. Mullerian Inhibiting Substance is an ovarian growth factor of emerging clinical significance. Fertil Steril 2007;88:539-46.


36. Gibreel A, Maheshwari A, Bhattacharya S, Johnson NP. Ultrasound tests of ovarian reserve: a systematic review of accuracy in predicting fertility outcomes. Hum Fertil (Camb) 2009;12:95-106.


37. Tufekci KU, Meuwissen RL, Genc S. The role of microRNAs in biological processes. Methods Mol Biol 2014;1107:15-31.


38. Xie S, Batnasan E, Zhang Q, Li Y. MicroRNA expression is altered in granulosa cells of ovarian hyperresponders. Reprod Sci 2016;23:1001-10.


39. Hong L, Peng S, Li Y, Fang Y, Wang Q, Klausen C, et al. MiR-106a increases granulosa cell viability and is downregulated in women with diminished ovarian reserve. J Clin Endocrinol Metab 2018;103:2157-66.



40. Karakaya C, Guzeloglu-Kayisli O, Uyar A, Kallen AN, Babayev E, Bozkurt N, et al. Poor ovarian response in women undergoing in vitro fertilization is associated with altered microRNA expression in cumulus cells. Fertil Steril 2015;103:1469-76.



41. Simoni M, Nieschlag E, Gromoll J. Isoforms and single nucleotide polymorphisms of the FSH receptor gene: implications for human reproduction. Hum Reprod Update 2002;8:413-21.



42. Romereim SM, Summers AF, Pohlmeier WE, Zhang P, Hou X, Talbott HA, et al. Gene expression profiling of bovine ovarian follicular and luteal cells provides insight into cellular identities and functions. Mol Cell Endocrinol 2017;439:379-94.


43. Tian X, Fei Q, Du M, Zhu H, Ye J, Qian L, et al. MiR-130a-3p regulated TGF-β1-induced epithelial-mesenchymal transition depends on SMAD4 in EC-1 cells. Cancer Med 2019;8:1197-208.



44. Kong X, Zhang J, Li J, Shao J, Fang L. MiR-130a-3p inhibits migration and invasion by regulating RAB5B in human breast cancer stem cell-like cells. Biochem Biophys Res Commun 2018;501:486-93.


45. Chen X, Yue B, Zhang C, Qi M, Qiu J, Wang Y, et al. MiR-130a-3p inhibits the viability, proliferation, invasion, and cell cycle, and promotes apoptosis of nasopharyngeal carcinoma cells by suppressing BACH2 expression. Biosci Rep 2017;37:BSR20160576.




46. Ostadrahimi S, Abedi Valugerdi M, Hassan M, Haddad G, Fayaz S, Parvizhamidi M, et al. MiR-1266-5p and miR-185-5p promote cell apoptosis in human prostate cancer cell lines. Asian Pac J Cancer Prev 2018;19:2305-11.


47. Ni W, Xia Y, Bi Y, Wen F, Hu D, Luo L. FoxD2-AS1 promotes glioma progression by regulating miR-185-5P/HMGA2 axis and PI3K/AKT signaling pathway. Aging (Albany NY) 2019;11:1427-39.



48. Shen F, Chang H, Gao G, Zhang B, Li X, Jin B. Long noncoding RNA FOXD2-AS1 promotes glioma malignancy and tumorigenesis via targeting miR-185-5p/CCND2 axis. J Cell Biochem 2019;120:9324-36.


49. Li W, Liang J, Zhang Z, Lou H, Zhao L, Xu Y, et al. MicroRNA-329-3p targets MAPK1 to suppress cell proliferation, migration and invasion in cervical cancer. Oncol Rep 2017;37:2743-50.


50. Lu G, Wong MS, Xiong MZ, Leung CK, Su XW, Zhou JY, et al. Circulating microRNAs in delayed cerebral infarction after aneurysmal subarachnoid hemorrhage. J Am Heart Assoc 2017;6:e005363.



51. He XM, Zheng YQ, Liu SZ, Liu Y, He YZ, Zhou XY. Altered plasma microRNAs as novel biomarkers for arteriosclerosis obliterans. J Atheroscler Thromb 2016;23:196-206.


52. Wang X, He X, Deng X, He Y, Zhou X. Roles of miR-4463 in H2O2-induced oxidative stress in human umbilical vein endothelial cells. Mol Med Rep 2017;16:3242-52.



53. He X, Du C, Zou Y, Long Y, Huang C, Chen F, et al. Downregulation of microRNA-4463 attenuates high-glucose- and hypoxia-induced endothelial cell injury by targeting PNUTS. Cell Physiol Biochem 2018;49:2073-87.


54. Wang X, Du C, He X, Deng X, He Y, Zhou X. MiR-4463 inhibits the migration of human aortic smooth muscle cells by AMOT. Biosci Rep 2018;38:BSR20180150.




55. Ding CF, Chen WQ, Zhu YT, Bo YL, Hu HM, Zheng RH. Circulating microRNAs in patients with polycystic ovary syndrome. Hum Fertil (Camb) 2015;18:22-9.


- TOOLS







